16S-EZ Frequently Asked Questions
Sample Preparation
What sample types can be submitted for 16S-EZ?
Genomic DNA (normalized to 20 ng/µL), fresh frozen tissue, or frozen cell pellets. For other sample types (e.g. soil or stool), please submit a request through our custom form.
Does GENEWIZ perform QC on my DNA samples prior to library preparation?
No, we do not perform sample QC in an effort to provide the fastest and most cost-effective service possible. Please measure the concentration of your samples (preferably using a dsDNA quantification assay such as Qubit or PicoGreen) and normalize to 20 ng/µL prior to submission.
Do DNA samples need to be normalized to 20 ng/µL?
Yes, GENEWIZ does not adjust the concentration prior to library preparation. Instead, we use equal volumes for each sample. For optimal results, please adjust your sample concentrations to 20 ng/µL prior to submission. We recommend using a dsDNA quantification assay such as Qubit® or PicoGreen™.
What is the 16S-EZ workflow?
The 16S-EZ workflow includes genomic DNA extraction (if necessary), amplification of the V3 and V4 hypervariable regions of 16S rRNA, library construction, sequencing on the Illumina® with 2x250 bp configuration, and data analysis.
What is the sensitivity of your 16S-EZ assay?
Our targeted 16S-EZ assay can typically detect down to genus level. Sensitivity is ultimately dependent on the classification of the organism in the data analysis database and the abundance of the taxa in the sample itself.
How does 16S-EZ compare to other 16S methods on the market?
In addition to industry-leading turnaround time, 16S-EZ uses a proprietary primer pool to target the hypervariable V3 and V4 regions of the 16S rRNA gene, which has shown to be more sensitive and accurate compared to V4-only sequencing.
Does GENEWIZ offer whole genome shotgun metagenomics?
Yes, please submit your request through the Whole Genome Metagenomics form.
What happens if my samples do not meet your starting material requirements?
Please contact us by submitting an inquiry here
Sample Submission
How should I prepare the tubes?
Prepare samples in microcentrifuge tubes. Label each tube clearly with the sample name, as it appears on the order form. See example below.

How can I send my samples to GENEWIZ?
You can send your samples, along with a copy of the order receipt, to GENEWIZ using one of the following options.
Option #1 (recommended): Ship samples directly to our facility using the shipping address listed on the order receipt. For DNA samples, we recommend shipping overnight on dry ice. However, you can ship at room temperature or on blue/dry ice. For cell pellets or frozen tissue, please ship overnight on dry ice.
Option #2: Submit samples via a local dropbox. We have an extensive network of convenient dropboxes throughout the US. Please submit an inquiry to find a dropbox near you. DNA samples may be deposited at room temperature. Please place the samples and order receipt together in a resealable plastic bag and contact us to schedule a pickup.
Results
How much data can I expect per sample?
GENEWIZ typically provides ~50,000 reads per sample.
What is the turnaround time?
Turnaround is 15 business days for purified DNA samples and 20-25 business days for frozen cell pellets or fresh frozen tissue.
What does the standard 16S-EZ data analysis include?
See below. Note that figures are for illustrative purposes only.
- Taxonomic classification
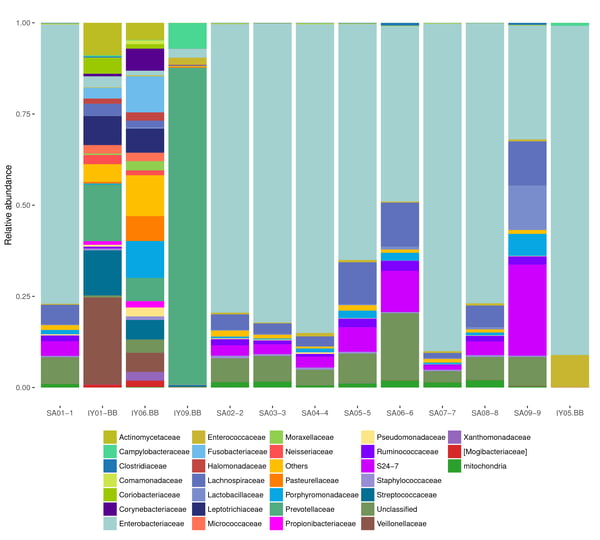
- Rank-abundance curve
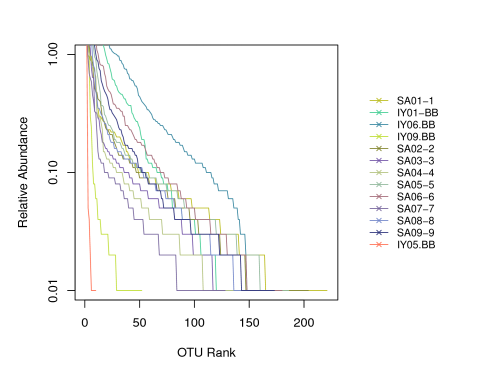
- Rarefaction curve
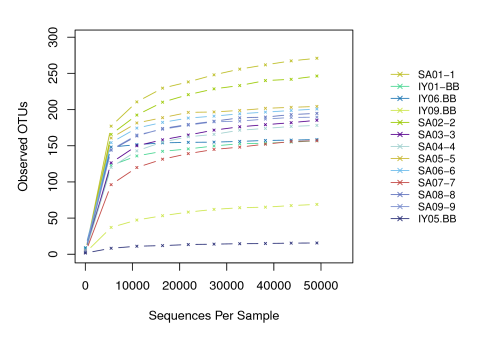
- Principal coordinate analysis
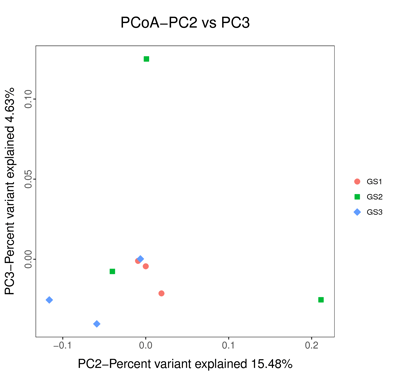
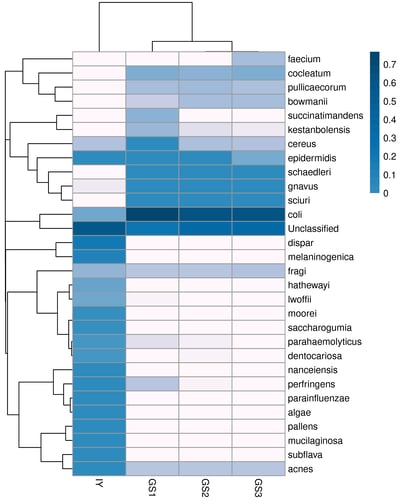
- Hierarchical clustering
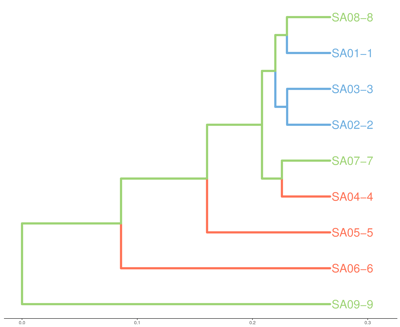
Will raw data be delivered to me?
Yes, all raw data (i.e. FASTQ files) will be delivered upon project completion.
How do I download my data?
Once the data analysis is complete, if applicable, a member of our bioinformatics team will reach out to you with a link to download your data from our sFTP server. Login credentials and instructions on how to retrieve your data will be included in the data delivery email. Please feel free to contact us should you have trouble retrieving your data. You can also refer to GENEWIZ's sFTP Data Download Guide for step-by-step instructions and troubleshooting tips.
Will my 16S-EZ data be secure at GENEWIZ?
We take data security very seriously and make every attempt to keep your data private and protected. The data transfer we offer is secure; however, should you desire an alternative delivery method, we are happy to work with you.
Have a specific question?
Contact Us | Phone 1-877-GENEWIZ (436-3949), ext. 1