Jump to Topic
Assay FAQ:
GENEWIZ FAQ:
What is single-cell RNA sequencing (RNA-Seq)?
Single-cell RNA-Seq provides transcriptional profiling of thousands of individual cells. This level of throughput analysis enables researchers to understand at the single-cell level what genes are expressed, in what quantities, and how they differ across thousands of cells within a heterogeneous sample.
The workflow for single-cell RNA-Seq is outlined below. Isolated cells are frozen by the customer according to GENEWIZ cryopreservation protocol and shipped overnight. At GENEWIZ, cell viability is assessed and, if necessary, dead cell removal may be performed (see below for more information). Cells are then processed by the 10x Genomics® Chromium™ X Controller to create single-cell libraries. The libraries are then pooled and sequenced on the Illumina® platform.
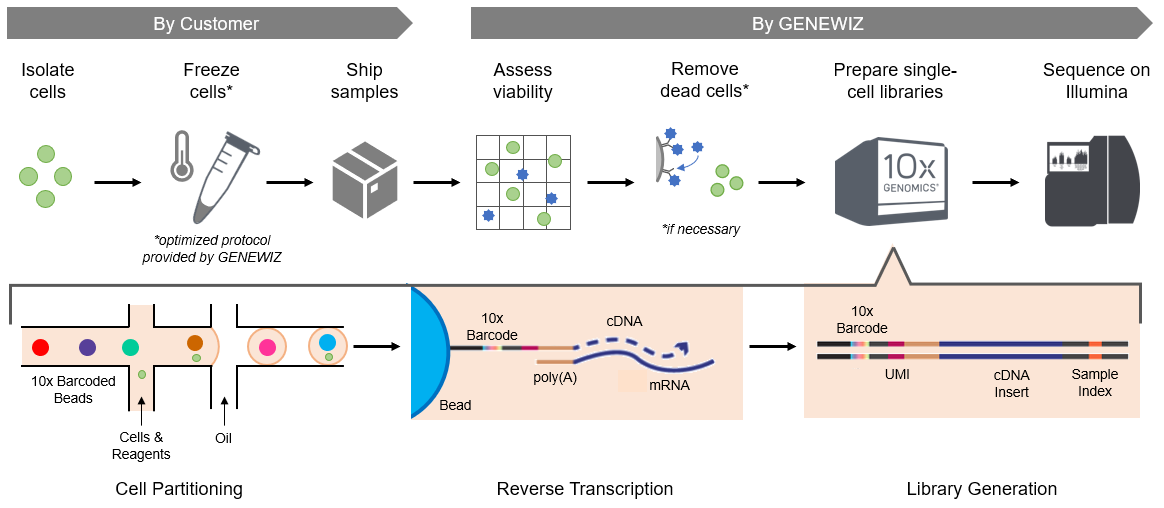
Within the Chromium X Controller, cells are loaded onto a microfluidic chip and encapsulated within droplets containing barcoded gel beads and reagents for reverse transcription. Following cell lysis, the beads capture the poly(A) tails of the mRNA molecules. Reverse transcription generates cDNA tagged with a 10x barcode to identify the cell and a unique molecular identifier (UMI) to label the mRNA transcript. The pooled cDNA is amplified in bulk, and Illumina adapters, including a sample index, are added to the fragments to generate sequencing libraries.
Read our article How Single-Cell Sequencing Works for more information.
How does single-cell RNA-Seq differ from standard and ultra-low input RNA-Seq?
Standard RNA-Seq produces a representative snapshot of the transcriptional state averaged across all cells. The caveat with traditional RNA-Seq is the resolution of individual cells and cellular subpopulations are lost. Single-cell RNA-Seq allows researchers to not only identify cellular subpopulations, but to fully interrogate them at the single-cell level within a heterogeneous sample.
Like standard RNA-Seq, ultra-low input RNA-Seq provides bulk expression analysis of the entire cell population; however, as the name implies, a very limited amount of starting material is used, as low as 10 pg or a few cells. Single-cell RNA-Seq requires at least 50,000 cells (1 million is recommended) as an input. See the Sample Preparation section below for more information about sample submission guidelines.
Read our article Top 3 Factors to Consider Before Starting a Single-Cell Sequencing Project to learn if single-cell sequencing is the right NGS approach for your experiment.
What is your sample processing workflow?
Our sample processing workflow is illustrated here.
Viability and cell counts are measured immediately upon sample receipt. To ensure the highest quality data, cells must be processed on the 10x Genomics® Chromium™ system as soon as possible. As such, we developed a predefined workflow when samples do not pass initial QC for viability or cell count. Please refer to the workflow linked above for more information.
Which single-cell assays do you offer?
In addition to standard 3’ and 5’ Single-Cell RNA-Seq, we offer Immuno-Seq, Feature Barcoding/CITE-Seq, Flex Fixed RNA, ATAC-Seq, and Multiome (RNA-Seq + ATAC-Seq). We also offer Regulated Single-Cell processing for all services listed in our GCP-compliant, CAP-accredited and CLIA-certified laboratory environment for clinical applications.
What is the price for single-cell RNA-Seq?
Please submit a quote request to receive accurate pricing information, as the cost depends on the details of the project.
.png?width=150&height=149&name=single-cell-rna-seq-clims%20(1).png)
You can also contact the NGS team by submitting an inquiry.
How do dead cells impact data quality?
Low cell viability causes unreliable cell recovery in the Chromium™ system and typically leads to missed sequencing targets and suboptimal results. Furthermore, ambient RNA released by dead or apoptotic cells increases background noise, compromising data quality. To maximize the amount of useful sequencing data in single-cell projects, the fraction of dead cells must be minimized. We recommend that cell viability exceeds 90%, with a minimum of 70%. Samples with less than 70% viability may undergo dead cell removal (DCR). A minimum of 106 cells and no previous treatment with magnetic beads is required for DCR.
Please refer to our workflow for more information.
How does dead cell removal work?
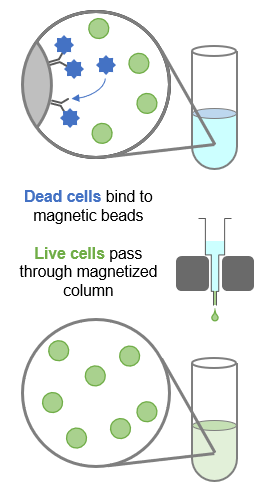
Dead cell removal (DCR) uses magnetic beads conjugated to antibodies against annexin V, a marker for dead and apoptotic cells (see image below). The beads capture dead or dying cells, enriching the sample for live cells.
How many cells can be sequenced per sample on the Chromium™ X Controller?
Revealing the full diversity of gene expression across a tissue or cell population often requires analysis of hundreds to thousands of viable cells. With our single-cell workflows, researchers have the flexibility to specify the number of cells to be sequenced, from 500 to 20,000 cells per sample. We typically recommend targeting 5,000-6,000 cells per sample for most experiments.
How many reads do I need for my experiment?
The number of reads required depends upon the genome size, the number of known genes, cell type, and transcripts. Generally, we recommend 50,000 reads per cell to maximize the identification of transcripts.
What starting materials are accepted?
We support a wide range of starting materials (including cryopreserved cells, isolated nuclei, fresh or fixed cells, and tissue samples) to meet the diverse needs of your single-cell research. For added convenience, we also offer optional nuclei isolation and tissue dissociation services for an additional fee.
For detailed sample submission requirements, please visit our sample submission guidelines.
What type of data analysis is available for single-cell RNA-Seq?
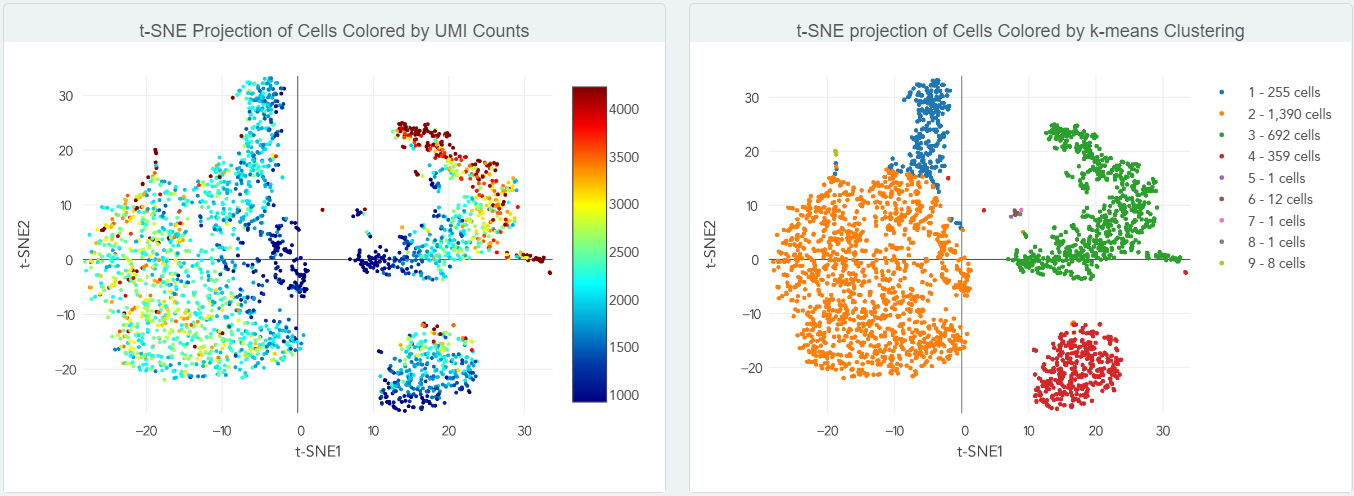
Here is a sample report. We use Cell Ranger software from 10x Genomics® for data analysis, which includes the following:
- Read alignment
- Feature-barcode matrix
- Digital gene expression matrix
- Clustering
- Gene expression analysis
- t-SNE projections (shown below)
How do I contact GENEWIZ for a technical consultation about my project?
The GENEWIZ NGS team is composed of Ph.D. scientists who can help you optimize your project design and provide consultation. You can contact the team by submitting an inquiry.
Does GENEWIZ guarantee turnaround?
Our team prioritizes fast turnaround times and provides a timeline based on first-pass processing. The turnaround time listed within the quote is inclusive of all steps quoted, unless otherwise noted. If repeat processing is required, the turnaround may be subject to change and will be proactively communicated to the client.
If the scope of a project changes after project initiation or if sample or project clarification is required after sample receipt, GENEWIZ may reassess the turnaround time based on the subsequent communications and modifications (if applicable).
What extraction methods are used?
All extraction kits and reagents are routinely updated to remain best-in-class. If you are seeking access to previous versions of our method, please submit an inquiry (availability is assessed on a case-by-case basis).
Extractions are performed to the best of our ability, and we are unable to guarantee yield due to multiple variables that may affect sample yield and quality. Costs to cover the work performed will be applied, regardless of the outcome.
What library preparation methods are used?
All preparation kits and reagents are routinely updated to remain best-in-class. If you are seeking access to previous versions of our method, please submit an inquiry (availability is assessed on a case-by-case basis).
Does GENEWIZ perform sample QC and what methods are used?
For most of the services, an initial sample QC is included within the fee of the project for all samples that will proceed to the next processing stage (e.g. library preparation or sequencing) unless otherwise noted within the quotation. Resubmissions or additional, optional samples may incur a nominal fee for QC.
Initial sample QC can include:
- Assessing RNA concentration and integrity
- Assessing DNA concentration and DNA size (for select projects)
- Assessing premade library size and concentration
- Assessing cryopreserved cells by cell count and viability
What sequencing instruments will be used for my project?
The sequencing platform will be listed within the Service Description of the quotation.
Unless specifically noted in the quotation, GENEWIZ reserves the right to choose between equivalent instruments depending on the target read depth and configuration requested. If a specific instrument is required, please add special comments in your quote/order and notify the GENEWIZ NGS team prior to project initiation.
You can contact the GENEWIZ NGS team at:
- US: NGS@azenta.com
- EU & UK: NGS.Europe@azenta.com
How does GENEWIZ guarantee data quality and yield?
Illumina-based projects
For samples that pass QC and libraries prepared at GENEWIZ:
Data Quality
- NovaSeq 2x150bp: ≥85% of bases ≥Q30
- NovaSeq 2x250bp: ≥80% of bases ≥Q30
- MiSeq 2x150bp: ≥80% of bases ≥Q30
- MiSeq 2x250bp: ≥75% of bases ≥Q30
Data Yield
- Within 10% of total data target yield per lane or flowcell, unless otherwise noted
- Within 20% of per sample target yield, unless otherwise noted
Quality and yield for samples that do not pass QC and are processed at best effort are not guaranteed. Premade libraries/library pools submitted for sequencing only quality and yield are evaluated on a case-by-case basis.
Multiplexing is performed to the best of our ability to ensure relatively even data distribution amongst samples.
PacBio-based projects
Due to various sample-related factors that may influence long-read sequencing yield and quality, GENEWIZ cannot guarantee overall data output and quality. However, based on extensive experience with long-read workflows, GENEWIZ has established target metrics based on the PacBio system based on the sample submitted and library type. If a project does not meet these targets, a thorough review of the processing will be performed. If the issue is determined to be unrelated to the sample, a repeat or top-off will be performed as necessary.
What is GENEWIZ’s coverage guarantee?
Due to the possible wide range of performance being influenced by sequence complexity and sample quality, GENEWIZ does not guarantee average coverage or on-target specificity for each sample. We can instead recommend target data output that would increase the chances of obtaining the desired coverage based on the sample and library type utilized.
What is GENEWIZ’s data analysis guarantee?
Data analysis is performed to the best of our ability and results are not guaranteed for samples or libraries not passing QC or do not fit the analysis pipeline’s criteria.
Can you recommend the best data delivery option?
Yes, we can recommend the best data delivery option based on the platform and project details. Options include:
- Secure File Transfer Protocol (sFTP) - additional charges may apply
- Customer Cloud Account - AWS, Microsoft Azure, Google Cloud
- Hard Drive - additional charges may apply
Delivery to most customer cloud solutions is free of charge. If you have any questions, please submit an inquiry.
How will my data be delivered?
By default, results are sent via a secure File Transfer Protocol (sFTP). Please refer to our sFTP Data Download Guide for instructions on how to download your data and troubleshooting tips. Additional charges may be applicable for large data transfers.
Please refer to the delivery email sent from GENEWIZ for detailed information on the transfer and consult with your IT department to ensure compliance with your institution’s policies.
Will my data be secure at GENEWIZ?
We take data security very seriously and we make every attempt to keep your data private and protected. The data transfer we offer is secure; however, should you desire an alternative delivery method, we are happy to work with you.
How long does GENEWIZ store samples?
We hold any remaining samples for up to 3 months after project completion. Clinical samples processed in our Regulatory-Environment or CLIA-licensed laboratory may have longer sample storage timelines. Please contact us if you would like samples retained for a longer period or shipped back to you.
How long does GENEWIZ store data?
GENEWIZ offers raw data storage (i.e. FASTQ for Illumina, BAM for PacBio, POD5 for Oxford Nanopore) for up to 6 months after project completion. Data generated for samples processed in the Regulatory-Environment or CLIA-licensed laboratory may have longer data retention timelines. Please contact us if you would like the data to be retained for a longer period.
How should I prepare and send my samples?
View our Sample Submission Guidelines for instructions on preparing and sending samples.
Ship samples directly to our facility. Use the shipping address listed on the order receipt.
What can I do if my samples do not meet the starting material requirements?
At GENEWIZ, we know that samples are valuable and not always perfect. We're here to help find solutions for working with less-than-ideal samples. Please reach out to us by submitting an inquiry, and we’ll work with you to meet your project needs
How do I request a quote?
For a quick tutorial, watch the video below.
How do I confirm a quote?
For a quick tutorial, watch the video below.
How can I monitor the progress of my project?
The status of your order, including an estimated date of delivery, can be viewed anytime through your GENEWIZ account. Visit the Order Summary page of your project to find the current order status.
How can I send my samples to GENEWIZ?
Option #1: Ship samples directly to our facility. Use the shipping address listed on your order receipt.
Option #2: For double-stranded DNA samples, submit samples into a local GENEWIZ dropbox which are conveniently located throughout the US, Europe and United Kingdom. You can easily locate a dropbox near you by logging into your account and choosing “Select from my dropboxes during the order checkout process. Place your order receipt and samples in a Ziploc bag packaged according to sample submission guidelines, and place in dropbox for pickup. If you do not have a dropbox near you, please request a new dropbox location here.
- Note: Not recommended for single-cell sequencing.
Have a specific question?
Contact Us | Phone 1-877-GENEWIZ (436-3949), ext. 1