Sample Preparation
What types of samples should I prepare for Plasmid-EZ?
Please submit purified plasmid DNA at a concentration of 20-200 ng/μL in a minimum volume of 10 μL in water or elution buffer. We also recommend a 260/230 ratio of 2.0-2.2 and 260/280 ratio of 1.8-2.0.
Samples should be prepared in microcentrifuge tubes, sealed 96-well plates, or capped PCR strip tubes. We will perform linearization upon receipt of plasmid DNA samples followed by library preparation and sequencing.
Should I use Nanodrop or Qbit to measure the concentration?
We recommend using Qubit as NanoDrop® cannot distinguish between dsDNA and ssDNA (oligos and dNTPs) and may cause you to overestimate the amount of dsDNA in your samples.
How should I prepare the tubes/plates?
Tubes for <48 Samples
If you are submitting <48 samples, please use 8-strip, 0.2 ml PCR tubes and caps. You can cut off empty tubes and use them next time.
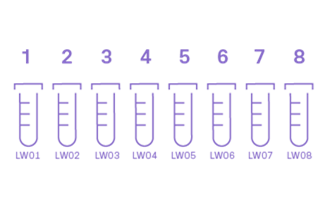
GENEWIZ's Online Ordering System will assign tube ID codes using your initials and the sample number (see Figure to the right). Please write these codes on the sides of your tubes using an indelible marker.
Label the side of your tubes with YOUR initials and sample number. These should match your order receipt.
Plates for >48 Samples
If you are submitting 48 or more samples, we recommend using a 96-well, semi-skirted PCR plate. Cap the wells with 8-strip caps. These are usually ordered separately from the plates. Be sure that the caps seal tightly! The semi-skirted plate helps to prevent the plate from bending in transit, resulting in fewer loose caps. Please ship the plate with cushioning to avoid potential damage.
Please contact us for help finding a vendor if your laboratory does not have any 96-well PCR plates or matching strip caps.
Arrange your samples vertically or horizontally. Please ensure that the plate orientation matches the order form well orientation.
What is the size range of plasmids that I can prepare for sequencing?
Plasmids up to 25 kb can be sequenced using this workflow. If you are sequencing larger than 25 kb, please contact DNAseq@azenta.com for a technical consultation.
Can a mixture of plasmids be submitted for sequencing using this service?
This service is intended for a clonal population of plasmids. If plasmid pools or mixtures are submitted, the sequencing and analysis results cannot be guaranteed.
What happens if my samples do not meet your starting material requirements?
Please contact DNAseq@azenta.com for technical consultation.
Sample Submission
When should I submit samples?
Samples may be submitted at any time. Samples must be received in the morning at our facility to be eligible for that day's cycle. Projects with incomplete information may be placed in the next sequencing cycle. Results are delivered on the same day in select regions or in as fast as 1 business day.
What are the sample submission guidelines for Plasmid-EZ?
Sample Type: Purified Plasmid DNA
Size: Up to 25 kb; if larger than 25 kb, please reach out to DNAseq@azenta.com.
Concentration: 20-200ng/μL in a minimum volume of 10μL. We recommend performing DNA quantification assay with Qubit or equivalent fluorometric method such as a plate reader (note: spectrophotometric methods like Nanodrop are much less accurate!).
Purity (A260/280): 1.8-2.0 and (260/230): 2.0-2.2
Buffer: Water, EB, or low TE (<0.1 mM EDTA)
How can I send my samples to GENEWIZ?
Please send your samples, along with a copy of your order receipt, to us using one of the following options.
Option #1: Submit samples into a local dropbox at room temperature. We have an extensive network of convenient dropboxes located throughout the US. To find a dropbox near you, please contact DNAseq@azenta.com.
Option #2: Ship samples directly to our NJ facility using the shipping address listed on your order receipt. You can ship at room temperature or on blue/dry ice.
What are the cutoff times for my dropbox pickup?
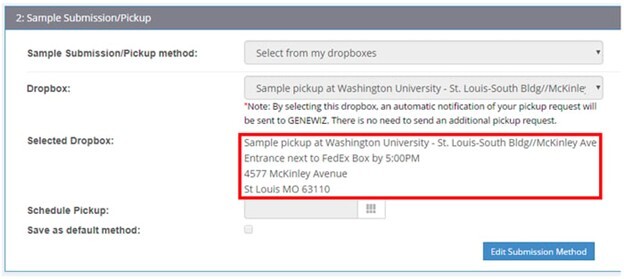
When submitting an order in our customer portal, the final step is to locate a dropbox closest to your institution; please choose the dropbox you will leave your sample at for Plasmid-EZ. During checkout, consult the Order Summary page to find out the daily cutoff times for your selected dropbox (see example below).
Ordering & Processing
How do I order Plasmid-EZ?
You can order Plasmid-EZ in minutes without a quote. Check out the Plasmid-EZ Order Guide for the 9 easy steps.
How much sequencing coverage is expected per plasmid?
Coverage is highly dependent on sample quality as determined by the end-user prior to sample submission and therefore cannot be guaranteed. If you have any questions or need troubleshooting assistance, please contact DNAseq@azenta.com.
How can I monitor the progress of my Plasmid-EZ project?
The estimated delivery date for your order can be viewed anytime through our customer portal on the Order Summary page. Because of the fast turnaround time of the service, we do not provide detailed project updates (i.e., timestamps for QC, library preparation, sequencing, and data analysis).
What is the turnaround time?
For customers located near one of our six overnight sequencing labs, you will receive results by 8 AM the next morning.
For customers who ship samples to our New Jersey facility, results will be available by the evening of the day that the samples are received.
Does GENEWIZ perform QC on my Plasmid-EZ samples prior to library preparation?
No, we do not perform sample QC. This enables us to provide rapid and cost-effective service for whole plasmid sequencing. Prior to submission, please measure the concentration of your samples (preferably using a dsDNA quantification assay such as Qubit or PicoGreen) and submit samples within a range of 20-200ng/μL.
Can GENEWIZ perform plasmid DNA preparation for me?
Plasmid preparation can be ordered after the Plasmid-EZ results are available. We will retain your submitted plasmid DNA samples for downstream plasmid preparation.
Alternatively, you can submit plasmid preparation order and choose to do Plasmid-EZ as sequence verification for your plasmid.
Results
Can Plasmid-EZ detect plasmid multimers?
Sometimes, multiple plasmid copies combine via homologous recombination to produce a multimeric form of plasmid with repeating sequences. This process is called plasmid multimerization. Plasmid-EZ is an effective way to detect such multimers; it generates exceptionally long reads that can detect large repeat sequences within a plasmid. However, if there are multimers that are present at a higher proportion in the sample, they can cause misassembly.
What data output will I receive for Plasmid-EZ?
You will receive an interactive data report, consensus sequence of your plasmid(s) in FASTA format, raw reads in FASTQ files, GenBank files, ab1 files, variation files, and annotation files, including plasmid map images. If a reference sequence is provided, then we will also provide an alignment file and variation files highlighting any differences between the consensus sequence and reference sequence.
How do I download my data?
Once data analysis is complete, you will receive an email that the order is completed. Log in to your GENEWIZ account. Click on “View results” tab of the order and download the results. The results will be downloaded as zipped files.
Is there additional bioinformatics available?
All custom bioinformatic analysis requests are taken on a case-by-case basis after a technical consultation with one of our technical support scientists. Please contact DNAseq@azenta.com for questions.
Troubleshooting
What if my plasmid samples fail to assemble?
It is likely that sequencing failure, i.e., failure of the sample to produce consensus sequence with 10x coverage or less, may be due to samples not prepared at the required DNA concentration, containing a mixed plasmid population, or degraded/fragmented DNA.
To achieve optimal sequencing results, please follow our recommende sample submission guidelines.
What if my results do not match my expectations or hypothesis?
We cannot guarantee the results of the sequencing data. If you feel that a processing error may have occurred, please contact DNAseq@azenta.com and we will investigate.
Colony-EZ
What is Colony-EZ?
Colony-EZ is our whole plasmid sequencing service, starting from bacteria instead of purified plasmid. We will isolate your plasmid preparation prior to sequencing for an added charge.
How do I submit my bacteria?
You can submit your bacteria in several different ways!
- Bacterial colonies: Plate colonies on agar plates. Circle and number colonies or leave blank for random pick. Wrap agar plates in parafilm and provide a cushion to protect against potential damage. If the colonies are too small or not visible, we will contact you and incubate the plate overnight at 37°C before processing, resulting in a delay of one day.
- Liquid culture: 100μL of overnight-grown liquid culture (8 to 16 hours), shipped in 200μL strip tubes. If the culture is not dense, we will contact you and grow the culture overnight at 37 °C before processing, resulting in a delay of one day.
- Colonies picked in water: Pick 1 large colony in 10μL H2O only (no media). Samples should be visibly cloudy. Please note, if the plasmid is low copy, submitting colonies picked in H2O might fail sequencing.
- Glycerol stock: The sample submitted should be cloudy. If the liquid is not cloudy, we will contact you and will incubate the sample overnight at 37°C before processing, resulting in a delay of one day.
Please visit our sample submission guidelines to learn more. Samples submitted outside our guidelines can result in sequencing failure and poor-quality results. We highly recommend following these guidelines. Note: High-copy plasmids are preferred (less than 15kb). Submit colonies that will grow in 37°C. Any cells that require modified growth conditions may not sequence well.
For low copy plasmids, please use our Colony-EZ plus service.
When will I receive the results?
For customers located near one of our six overnight sequencing labs, you will receive results by 8am the next morning.
For customers who ship samples to our New Jersey facility, results will be available by the evening of the day that the samples are received.
What is Colony-EZ plus?
Colony-EZ plus is our whole plasmid sequencing service, starting from bacteria instead of purified plasmid. In this service, we do crude plasmid prep and then perform sequencing.
What is the difference between Colony-EZ and Colony-EZ plus service?
| Colony-EZ | Colony-EZ Plus | |
| Description | We sequence directly from the colonies | We grow the cells overnight, do crude plasmid prep and then sequence using whole plasmid sequencing workflow |
| TAT | Next day by 8 AM in most locations | Adds extra 1–2-day TAT |
| Price | $19.99/sample | $24.99/sample |
| Sample Type Accepted |
|
|
| When To Choose? | For high copy plasmids, and not difficult plasmids. | Low copy plasmids; difficult plasmids; for deeper coverage |
Will the plasmid preparation used for whole plasmid sequencing be returned?
The plasmid preparation used is an unpure prep and may not be suitable for downstream processes; therefore, plasmid preparations are not returned for this service.
Have a specific question?
Email: dnaseq@azenta.com | Phone: 1-877-GENEWIZ (436-3949), ext. 2